Warning
This documents an unmaintained version of NetworkX. Please upgrade to a maintained version and see the current NetworkX documentation.
cn_soundarajan_hopcroft¶
-
cn_soundarajan_hopcroft(G, ebunch=None, community='community')[source]¶ - Count the number of common neighbors of all node pairs in ebunch
- using community information.
For two nodes
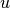 and
and  , this function computes the number of
common neighbors and bonus one for each common neighbor belonging to
the same community as
, this function computes the number of
common neighbors and bonus one for each common neighbor belonging to
the same community as 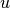 and
and  . Mathematically,
. Mathematically,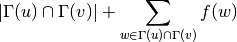
where
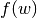 equals 1 if
equals 1 if 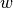 belongs to the same community as
belongs to the same community as 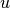 and
and  or 0 otherwise and
or 0 otherwise and 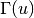 denotes the set of
neighbors of
denotes the set of
neighbors of 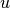 .
.Parameters: - G (graph) – A NetworkX undirected graph.
- ebunch (iterable of node pairs, optional (default = None)) – The score will be computed for each pair of nodes given in the iterable. The pairs must be given as 2-tuples (u, v) where u and v are nodes in the graph. If ebunch is None then all non-existent edges in the graph will be used. Default value: None.
- community (string, optional (default = ‘community’)) – Nodes attribute name containing the community information. G[u][community] identifies which community u belongs to. Each node belongs to at most one community. Default value: ‘community’.
Returns: piter – An iterator of 3-tuples in the form (u, v, p) where (u, v) is a pair of nodes and p is their score.
Return type: iterator
Examples
>>> import networkx as nx >>> G = nx.path_graph(3) >>> G.node[0]['community'] = 0 >>> G.node[1]['community'] = 0 >>> G.node[2]['community'] = 0 >>> preds = nx.cn_soundarajan_hopcroft(G, [(0, 2)]) >>> for u, v, p in preds: ... '(%d, %d) -> %d' % (u, v, p) ... '(0, 2) -> 2'
References
[1] Sucheta Soundarajan and John Hopcroft. Using community information to improve the precision of link prediction methods. In Proceedings of the 21st international conference companion on World Wide Web (WWW ‘12 Companion). ACM, New York, NY, USA, 607-608. http://doi.acm.org/10.1145/2187980.2188150