Warning
This documents an unmaintained version of NetworkX. Please upgrade to a maintained version and see the current NetworkX documentation.
jaccard_coefficient¶
-
jaccard_coefficient(G, ebunch=None)[source]¶ Compute the Jaccard coefficient of all node pairs in ebunch.
Jaccard coefficient of nodes
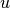 and
and  is defined as
is defined as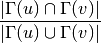
where
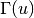 denotes the set of neighbors of
denotes the set of neighbors of 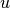 .
.Parameters: - G (graph) – A NetworkX undirected graph.
- ebunch (iterable of node pairs, optional (default = None)) – Jaccard coefficient will be computed for each pair of nodes given in the iterable. The pairs must be given as 2-tuples (u, v) where u and v are nodes in the graph. If ebunch is None then all non-existent edges in the graph will be used. Default value: None.
Returns: piter – An iterator of 3-tuples in the form (u, v, p) where (u, v) is a pair of nodes and p is their Jaccard coefficient.
Return type: iterator
Examples
>>> import networkx as nx >>> G = nx.complete_graph(5) >>> preds = nx.jaccard_coefficient(G, [(0, 1), (2, 3)]) >>> for u, v, p in preds: ... '(%d, %d) -> %.8f' % (u, v, p) ... '(0, 1) -> 0.60000000' '(2, 3) -> 0.60000000'
References
[1] D. Liben-Nowell, J. Kleinberg. The Link Prediction Problem for Social Networks (2004). http://www.cs.cornell.edu/home/kleinber/link-pred.pdf